Ah, Malignant Hyperthermia…always worry about it, practice how to treat it (OPEN your PediCrisis v 2 app!), and hopefully never see it or need to treat it. Since most cases of MH are associated with variants in the gene encoding the skeletal muscle ryanodine receptor 1, RYR1, could next-generation gene sequencing help us identify patients at risk? As many of you know, Ron Litman of blessed memory started the Pediatric Anesthesia Article of the Day and was deeply involved with the Malignant Hyperthermia Association of the U.S. (MHAUS). Thus, I thought this would be a perfect article to honor his memory. Admittedly though, this is definitely NOT my area of expertise, so I asked Jerry Parness, who definitely is an expert to review this article for us. Myron Yaster MD
Original article
van den Bersselaar LR, Hellblom A, Gashi M, Kamsteeg EJ, Voermans NC, Jungbluth H, de Puydt J, Heytens L, Riazi S, Snoeck MMJ.Referral. Indications for Malignant Hyperthermia Susceptibility Diagnostics in Patients without Adverse Anesthetic Events in the Era of Next-generation Sequencing. Anesthesiology. 2022 Jun 1;136(6):940-953. PMID: 35285867
A bane of our existence as anesthesiologists has always been meeting our patients and families in the preop area and being told that some relative in the past three generations either died or survived an episode of supposed Malignant Hyperthermia (MH), and that they were told by their doctors to always tell this to the anesthesiologist. Did they get tested for MH? NO! Did any direct descendent get tested for MH in any manner, shape or form? NO! So, how are we to move ahead? Should we practice CYA (or CYB, Cover Your Bum, for polite company) medicine and do a non-triggering anesthetic, or just move ahead with a standard anesthetic, but expectantly, with dantrolene and your pedicrisis app at the ready, nearby?
The vast majority of MH causing mutations are in RyR1, the skeletal muscle Ca2+ channel responsible for the Ca2+ release in excitation-contraction coupling (ECC). Some, actually very few patients have mutations in CACNA1S, the alpha-1 subunit of the skeletal muscle dihydropyridine receptor, the voltage change sensor responsible for triggering excitation-contraction coupling by a direct interaction with RyR1, and STAC3, the SH3 and cysteine-rich domain 3 protein, a muscle protein that is involved in modulating Ca2+ release in ECC, mutations in which are responsible for Native American myopathy and MH susceptibility (MHS). Given the data we see below, there must be other proteins(?), mutations which are responsible for MHS.
Clearly without easy, affordable, and reliable diagnostic tests in our armamentarium, there is no easy answer on how to proceed clinically in these incompletely described situations. We see this even more so from today’s PAAD by van den Bersselaar et al. The paper agglomerates data from four MH diagnostic centers (Lund, Sweden; Antwerp, Belgium; Nijmegen, The Netherlands; and Toronto, Ontario, Canada) from the years 2010 to 2019. They examined their data for patients with with a personal or family history of adverse anesthetic events or MH, those with RyR1 or other myopathies, those with history of elevated creatine kinase (CPK), and those with a history of exertion heat stroke. Moreover, and for the more scientifically inclined of our colleagues, they examined all unclassified RyR1 missense variants by a recently described and fairly well validated computational score called REVEL (Rare Exome Variant Ensemble Learner), which uses multiple, different, previously described, computational protein amino acid pathogenicity predictor programs in an ensemble program that actually performs better at predicting variant pathogenicity than any one of the programs that it uses in the ensemble (1). Using this score, if the RyR1, or any of the other MH-related gene variants are known, then one can hopefully predict the likelihood that such a variant can cause MH under volatile gas anesthesia, and help us proceed accordingly without having to go through laborious, expensive biological testing.
During the study period, the number of patients without a personal or family history of an adverse anesthetic event referred to these diagnostic units has increased. Nevertheless, nearly 40% of those were eventually diagnosed as malignant hyperthermia susceptible (MHS). That is a sobering thought. That number, however, cannot be interpreted as all comers to a preop unit have a 40% chance of being MHS. The patients referred to a MH diagnostic center are a biased sample, and include those with myopathies, a history of exertional heat stroke, elevated CPK levels, or other conditions that in the past have been associated, rightly or wrongly, with MH susceptibility.
What emerges in the data is striking. Of those with a personal history of suspected MH, fully 44% were eventually diagnosed as MHS, while about 20% were MHN and the rest unknown for incomplete data. With a positive family history (FH) of suspected MH, 40% were MHS, 39% MHN, and 30% unknown. Rhabdomyolysis referrals were MHS 41% of the time, MHN in ~22%, and unknown in 37% of referrals. Surprisingly, though the sample size was very small, only ~20% of those with a personally diagnosed RyR1 myopathy were MHS - meaning that not every RyR1 variant signifies MHS. If you had a family history of a RyR1 variant of unknown significance, only 10% tested as MHS. Your level of comfort in treating such patients with VAs will be based on some data and your psychological need to dispense CYB care. “La plus ca change, la plus ca meme chose”!
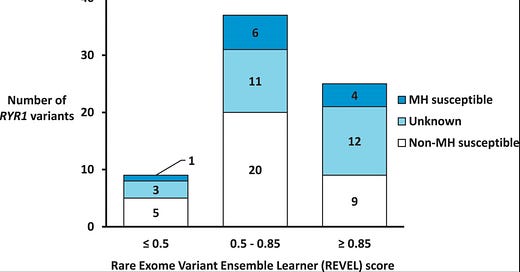
And now, for the strawberry shortcake/pumpkin pie/ice cream sundae with all the works AND whip cream…. we must have whip cream! For dessert, we must know how REVEL helps us in determining whether a particular, non-curated variant in RyR1, CACNA1,or STAC3, is pathogenic. The answer is to be found in their Figure 3 (below). The highest REVEL score (>/= 0.85) should predict pathogenicity to a high degree, but when the known MH status of study patients was examined by REVEL, 25 patients fell into this high score category, but only 4 were known MHS, 9 were known MHN, and 12 were in the unknown (insufficient data) category. As far as algorithmic diagnosis, we’ve got a long way to go, baby.
This is a paper worth going through, not for answers to our ultimate questions, but in order to understand how we, as a specialty try, and how hard it is to actually get actionable clinical data.
References
1. Ioannidis, NM, et al., REVEL: An ensemble method for predicting the pathogenicity of rare missense variants. Am J Hum Genet. 2016;99(4): 877-885. doi:10.1016/j.ajhg.2016.08.016.